Why normal cells turn into cancer cells? One of the factors is deeply related to the failure of the cell differentiation mechanism called DNA methylation (*1) . The joint research groups of The Institute of Medical Science, the University of Tokyo, Yokohama City University, and Center for Integrated Protein Science Munich (CIPSM) have clarified new mechanism for controlling DNA methylation in cells.
Ubiquitination(*2) of a protein called PAF15(*3) is an important factor for the inheritance of DNA methylation, according to the new research. The group also successfully demonstrated a molecular mechanism by which PAF15 is ubiquitinated. In this way, the group revealed the mechanism underlying how cellular memory is inherited when cells proliferate.
The research was published in Nature communications (online version, March 6, 2020). The results are expected to greatly contribute to applications in this field, such as the development of new inhibitors of DNA methyltransferase that specifically target cancer cells.
The primary mechanism of this DNA methylation has not been fully elucidated

© 2020 IMSUT, The University of Tokyo CC-BY
There are two factors that transform normal cells into cancer cells. One is a genetic mutation that alters the DNA sequence, and the other is an epigenetic mutation(*4) that changes how genes are used. An epigenetic mutation is defined as an aberrant pattern of “DNA methylation” and “histone modification” (*5) . When cells proliferate, both genetic and epigenetic mutations are transmitted to new cells.
The research team decided to investigate the mechanism of faithful "DNA methylation inheritance" which is critical for cancer suppression and has not been sufficiently investigated. As one of the lead researchers, associate professor Atsuya Nishiyama of the University of Tokyo, explains as follows:
"Every cell has the same genetic information. Each cell has very different characteristics. It is DNA methylation that determines the characteristics of each cell. Until now, the correlation between the failure of the mechanism of DNA methylation and cancer of cells was known, but how DNA methylation controls disease was not. That's why we conducted research to understand the basic mechanisms of DNA methylation. "
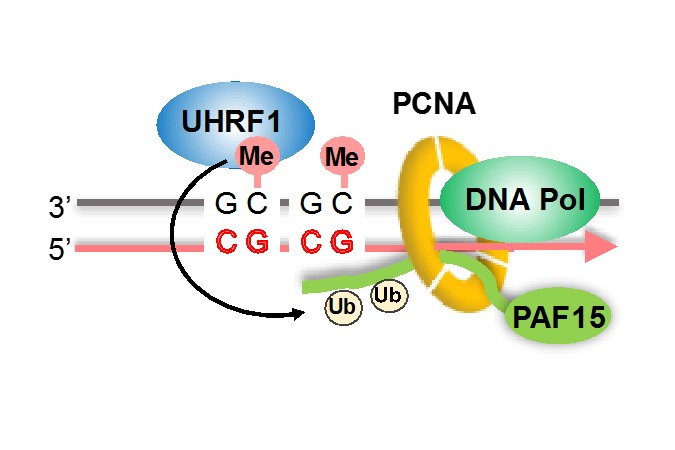
Further analysis revealed that PAF15 binds to chromosomes via PCNA(*7) during DNA replication, The research group also showed that two lysine residues of PAF15 undergo monoubiquitination by UHRF1(*8) to facilitate DNMT1 chromatin localization. These results indicate that PAF15 is an important regulator of DNMT1.
(*2) ubiquitination: the reaction that covalently conjugates a small polypeptide, ubiquitin, to a particular residue of a target protein. Ubiquitination affects protein stability, cellular localization, enzymatic activity, and protein-protein interactions.
(*3) PAF15: PCNA-associated factor 15 was identified as a proliferating nuclear antigen interacting protein.
(*4) epigenetic mutation: defined as aberrant DNA methylation and histone modifications.
(*5) histone modification: Covalent post-translational modification to histone proteins, which includes methylation, acetylation, and ubiquitination. These modifications cooperate with DNA methylation to impact gene expression and chromatin structure.
(*7) PCNA: Proliferating cell nuclear antigen (PCNA) is a critical regulator of DNA replication, which regulates DNA polymerase processivity.
(*8) UHRF1: Ubiquitin-like, PHD and RING finger containing 1 plays a critical role in the maintenance of DNA methylation. UHRF1 specifically binds to hemi-methylated DNA during S-phase and recruits DNMT1 to sites of DNA methylation.
http://www.ims.u-tokyo.ac.jp/imsut/en/
Yokohama City University
https://www-user.yokohama-cu.ac.jp/~english/
Center for Integrated Protein Science Munich (CIPSM)
https://www.en.uni-muenchen.de/index.html
PAF15 is a factor guaranteeing the maintenance of DNA methylation
The research group conducted experiments using a cell-free system derived from Xenopus laevis eggs to analyze the DNMT1 protein complex. Their work resulted in the new discovery of PAF15 as a factor that specifically binds to DNMT1(*6) .
Further analysis revealed that PAF15 binds to chromosomes via PCNA(*7) during DNA replication, The research group also showed that two lysine residues of PAF15 undergo monoubiquitination by UHRF1(*8) to facilitate DNMT1 chromatin localization. These results indicate that PAF15 is an important regulator of DNMT1.
During unperturbed S-phase, most of the DNMT1 on the chromosome was bound to ubiquitinated PAF15. Elevated histone H3 ubiquitination levels and the interaction between DNMT1 and ubiquitinated H3 were observed in conjunction with the inhibition of PAF15 function.
This finding suggests that PAF15 ubiquitination is the primary pathway controlling DNMT1 localization to DNA methylation sites, and that histone H3 ubiquitination may serve as a backup system. The team also introduced mutations to amino acids at the ubiquitination site of PAF15 in mouse ES cells, and found that the level of DNA methylation in the whole genome was greatly reduced, thus making it clear that PAF15 was a factor guaranteeing the maintenance of DNA methylation.
※ Please see the research group's paper for details:
Potential to greatly contribute to the development of DNA methylase inhibitors
DNA methylase is also attracting attention as a material for anticancer drugs. The results of this study have shown not only the scientific significance of clarifying the new mechanism of DNA methylation inheritance, but also the potential to greatly contribute to the development of DNA methyltransferase inhibitors.
In addition, it has been reported that PAF15 is over-expressed in various cancer cells. It will be important to clarify the effect of PAF15 on DNA methylation control in the future.

PAF15 in a complex with PCNA undergoes UHRF1-mediated dual mono-ubiquitylation, which is essential for DNMT1 recruitment, and subsequent maintenance of DNA methylation.
Research Notes
(*1) DNA methylation: an epigenetic modification that occurs by the addition of a methyl (CH3) group to DNA, affecting gene expression and chromatin structure.
(*2) ubiquitination: the reaction that covalently conjugates a small polypeptide, ubiquitin, to a particular residue of a target protein. Ubiquitination affects protein stability, cellular localization, enzymatic activity, and protein-protein interactions.
(*3) PAF15: PCNA-associated factor 15 was identified as a proliferating nuclear antigen interacting protein.
(*4) epigenetic mutation: defined as aberrant DNA methylation and histone modifications.
(*5) histone modification: Covalent post-translational modification to histone proteins, which includes methylation, acetylation, and ubiquitination. These modifications cooperate with DNA methylation to impact gene expression and chromatin structure.
(*6) DNMT1: DNA methyltransferase 1 (DNMT1) is an enzyme that catalyzes the transfer of methyl groups to cytosine. DNMT1 is responsible for maintenance DNA methylation which ensures the inheritance of DNA methylation in proliferating cells.
(*7) PCNA: Proliferating cell nuclear antigen (PCNA) is a critical regulator of DNA replication, which regulates DNA polymerase processivity.
(*8) UHRF1: Ubiquitin-like, PHD and RING finger containing 1 plays a critical role in the maintenance of DNA methylation. UHRF1 specifically binds to hemi-methylated DNA during S-phase and recruits DNMT1 to sites of DNA methylation.
About the research
Atsuya Nishiyama*, Christopher B. Mulholland, Sebastian Bultmann, Satomi Kori, Akinori Endo, Yasushi Saeki, Weihua Qin, Carina Trummer, Yoshie Chiba, Haruka Yokoyama, Soichiro Kumamoto, Toru Kawakami, Hironobu Hojo, Genta Nagae, Hiroyuki Aburatani, Keiji Tanaka, Kyohei Arita*, Heinrich Leonhardt* and Makoto Nakanishi*(*corresponding author),2020,“Two distinct modes of DNMT1 recruitment ensure stable maintenance DNA methylation”Nature communications.
Papers
Article URL: https://www.nature.com/articles/s41467-020-15006-4Related Links
The Institute of Medical Science, The University of Tokyohttp://www.ims.u-tokyo.ac.jp/imsut/en/
Yokohama City University
https://www-user.yokohama-cu.ac.jp/~english/
Center for Integrated Protein Science Munich (CIPSM)
https://www.en.uni-muenchen.de/index.html